What Does Condensed Dna Makeup A Chromosome
The most of import office of Deoxyribonucleic acid is to carry genes, the data that specifies all the proteins that make up an organism—including information most when, in what types of cells, and in what quantity each poly peptide is to be fabricated. The genomes of eucaryotes are divided upwards into chromosomes, and in this department nosotros see how genes are typically bundled on each chromosome. In addition, we describe the specialized DNA sequences that allow a chromosome to exist accurately duplicated and passed on from one generation to the next.
Nosotros also confront the serious challenge of DNA packaging. Each man cell contains approximately two meters of DNA if stretched finish-to-end; yet the nucleus of a human being jail cell, which contains the Dna, is only about half dozen μm in diameter. This is geometrically equivalent to packing 40 km (24 miles) of extremely fine thread into a tennis ball! The circuitous task of packaging DNA is achieved by specialized proteins that bind to and fold the DNA, generating a series of coils and loops that provide increasingly college levels of organization, preventing the Dna from becoming an unmanageable tangle. Amazingly, although the DNA is very tightly folded, information technology is compacted in a way that allows it to hands get bachelor to the many enzymes in the cell that replicate information technology, repair it, and use its genes to produce proteins.
Eucaryotic Deoxyribonucleic acid Is Packaged into a Set of Chromosomes
In eucaryotes, the Deoxyribonucleic acid in the nucleus is divided betwixt a fix of different chromosomes. For instance, the human being genome—approximately 3.2 × 109 nucleotides—is distributed over 24 dissimilar chromosomes. Each chromosome consists of a single, enormously long linear DNA molecule associated with proteins that fold and pack the fine Dna thread into a more compact structure. The complex of Dna and protein is called chromatin (from the Greek chroma, "color," because of its staining backdrop). In improver to the proteins involved in packaging the Dna, chromosomes are besides associated with many proteins required for the processes of factor expression, Dna replication, and Dna repair.
Bacteria comport their genes on a single DNA molecule, which is commonly round (meet Figure 1-thirty). This Dna is associated with proteins that package and condense the DNA, but they are different from the proteins that perform these functions in eucaryotes. Although often called the bacterial "chromosome," it does not accept the aforementioned construction as eucaryotic chromosomes, and less is known almost how the bacterial Deoxyribonucleic acid is packaged. Even less is known well-nigh how Dna is compacted in archaea. Therefore, our discussion of chromosome structure will focus nigh entirely on eucaryotic chromosomes.
With the exception of the germ cells, and a few highly specialized jail cell types that cannot multiply and lack DNA altogether (for example, red claret cells), each human being cell contains 2 copies of each chromosome, one inherited from the female parent and one from the father. The maternal and paternal chromosomes of a pair are called homologous chromosomes (homologs). The only nonhomologous chromosome pairs are the sex chromosomes in males, where a Y chromosome is inherited from the father and an X chromosome from the mother. Thus, each human cell contains a full of 46 chromosomes—22 pairs mutual to both males and females, plus ii and then-called sex chromosomes (X and Y in males, 2 Xs in females). DNA hybridization (described in detail in Chapter 8) tin be used to distinguish these human chromosomes by "painting" each one a dissimilar color (Figure 4-10). Chromosome painting is typically done at the stage in the cell cycle when chromosomes are especially compacted and easy to visualize (mitosis, run into beneath).

Effigy four-10
Man chromosomes. These chromosomes, from a male, were isolated from a jail cell undergoing nuclear partitioning (mitosis) and are therefore highly compacted. Each chromosome has been "painted" a dissimilar color to permit its unambiguous identification (more...)
Another more than traditional way to distinguish one chromosome from some other is to stain them with dyes that produce a striking and reliable pattern of bands along each mitotic chromosome (Figure 4-xi). The structural bases for these banding patterns are not well understood, and nosotros return to this event at the end of the affiliate. Withal, the design of bands on each type of chromosome is unique, allowing each chromosome to be identified and numbered.

Figure 4-11
The banding patterns of human being chromosomes. Chromosomes 1–22 are numbered in estimate social club of size. A typical human somatic (non-germ line) cell contains ii of each of these chromosomes, plus two sex activity chromosomes—two Ten chromosomes in (more than...)
The display of the 46 homo chromosomes at mitosis is called the human karyotype. If parts of chromosomes are lost, or switched between chromosomes, these changes can be detected by changes in the banding patterns or by changes in the pattern of chromosome painting (Figure 4-12). Cytogeneticists use these alterations to detect chromosome abnormalities that are associated with inherited defects or with sure types of cancer that arise through the rearrangement of chromosomes in somatic cells.

Figure 4-12
An aberrant man chromosome. (A) Two pairs of chromosomes, stained with Giemsa (see Figure iv-11), from a patient with clutter, a disease characterized by progressive deterioration of motor skills. The patient has a normal pair of chromosome 4s (left-hand (more than...)
Chromosomes Contain Long Strings of Genes
The most important function of chromosomes is to carry genes—the functional units of heredity. A gene is usually defined every bit a segment of Dna that contains the instructions for making a detail poly peptide (or a set of closely related proteins). Although this definition holds for the bulk of genes, several percent of genes produce an RNA molecule, instead of a protein, as their final product. Similar proteins, these RNA molecules perform a diverse set of structural and catalytic functions in the cell, and we talk over them in detail in subsequent capacity.
As might be expected, a correlation exists between the complexity of an organism and the number of genes in its genome (come across Table one-1). For example, total gene numbers range from less than 500 for simple bacteria to virtually 30,000 for humans. Bacteria and some single-celled eucaryotes have especially compact genomes; the complete nucleotide sequence of their genomes reveals that the DNA molecules that make up their chromosomes are little more than than strings of closely packed genes (Effigy four-13; see also Figure 1-30). Even so, chromosomes from many eucaryotes (including humans) contain, in improver to genes, a large excess of interspersed DNA that does not seem to carry critical information. Sometimes called junk Dna to signify that its usefulness to the cell has non been demonstrated, the particular nucleotide sequence of this DNA may not be important; simply the Dna itself, by interim as spacer material, may be crucial for the long-term evolution of the species and for the proper expression of genes. These issues are taken up in detail in Chapter 7.

Figure 4-13
The genome of South. cerevisiae (budding yeast). (A) The genome is distributed over sixteen chromosomes, and its complete nucleotide sequence was determined by a cooperative effort involving scientists working in many different locations, as indicated (grayness, Canada; (more...)
In general, the more than complex the organism, the larger its genome, but because of differences in the amount of excess DNA, the relationship is not systematic (come across Figure 1-38). For example, the human being genome is 200 times larger than that of the yeast S. cerevisiae, only 30 times smaller than that of some plants and amphibians and 200 times smaller than a species of amoeba. Moreover, because of differences in the amount of excess DNA, the genomes of similar organisms (bony fish, for case) can vary several hundredfold in their DNA content, fifty-fifty though they comprise roughly the same number of genes. Whatever the excess Dna may do, information technology seems clear that information technology is not a nifty handicap for a higher eucaryotic cell to deport a large amount of it.
The circulation of the genome over chromosomes also differs from ane eucaryotic species to the next. For instance, compared with 46 for humans, somatic cells from a species of small deer contain simply vi chromosomes, while those from a species of carp contain over 100. Even closely related species with similar genome sizes can have very different numbers and sizes of chromosomes (Figure 4-14). Thus, in that location is no uncomplicated relationship between chromosome number, species complexity, and total genome size. Rather, the genomes and chromosomes of modern-day species accept each been shaped by a unique history of seemingly random genetic events, acted on past selection pressures.

Effigy 4-14
Two closely related species of deer with very different chromosome numbers. In the development of the Indian muntjac, initially separate chromosomes fused, without having a major effect on the fauna. These ii species have roughly the same number of genes. (more than...)
The Nucleotide Sequence of the Man Genome Shows How Genes Are Arranged in Humans
When the Deoxyribonucleic acid sequence of human chromosome 22, 1 of the smallest human chromosomes (meet Figure four-11), was completed in 1999, information technology became possible for the first time to see exactly how genes are bundled forth an unabridged vertebrate chromosome (Figure 4-15 and Tabular array 4-ane). With the publication of the "first draft" of the unabridged human genome in 2001, the genetic landscape of all human chromosomes all of a sudden came into sharp focus. The sheer quantity of information provided by the Homo Genome Project is unprecedented in biology (Figure four-xvi and Table four-1); the human genome is 25 times larger than whatever other genome sequenced so far, and is 8 times as big as the sum of all previously sequenced genomes. At its peak, the Homo Genome Project generated raw nucleotide sequences at a rate of yard nucleotides per second around the clock. It will be many decades before this information is fully analyzed, but it volition proceed to stimulate many new experiments and has already afflicted the content of all the chapters in this book.

Figure 4-15
The organization of genes on a human chromosome. (A) Chromosome 22, one of the smallest homo chromosomes, contains 48 × ten6 nucleotide pairs and makes up approximately ane.5% of the unabridged man genome. About of the left arm of chromosome 22 consists (more...)
Table 4-1
Vital Statistics of Human Chromosome 22 and the Unabridged Man Genome.

Figure 4-sixteen
Scale of the human genome. If each nucleotide pair is drawn as ane mm as in (A), and so the human genome would extend 3200 km (approximately 2000 miles), far enough to stretch across the middle of Africa, the site of our human origins (red line in B). At (more...)
Although there are many aspects to analyzing the homo genome, here we merely make a few generalizations regarding the organisation of genes in human chromosomes. The beginning hitting feature of the human genome is how little of it (but a few percentage) codes for proteins or structural and catalytic RNAs (Figure 4-17). Much of the remaining chromosomal DNA is made up of curt, mobile pieces of DNA that have gradually inserted themselves in the chromosome over evolutionary time. We discuss these transposable elements in particular in later on chapters.
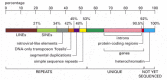
Figure 4-17
Representation of the nucleotide sequence content of the human being genome. LINES, SINES, retroviral-similar elements, and Deoxyribonucleic acid-only transposons are all mobile genetic elements that accept multiplied in our genome past replicating themselves and inserting the new (more...)
A second notable feature of the human genome is the large average factor size of 27,000 nucleotide pairs. As discussed above, a typical gene carries in its linear sequence of nucleotides the information for the linear sequence of the amino acids of a poly peptide. Only about 1300 nucleotide pairs are required to encode a protein of average size (about 430 amino acids in humans). Nearly of the remaining DNA in a gene consists of long stretches of noncoding Dna that interrupt the relatively short segments of DNA that code for poly peptide. The coding sequences are called exons; the intervening (noncoding) sequences are chosen introns (meet Figure 4-xv and Tabular array 4-1).
The majority of human genes thus consist of a long string of alternating exons and introns, with most of the gene consisting of introns. In dissimilarity, the majority of genes from organisms with meaty genomes lack introns. This accounts for the much smaller size of their genes (about one-twentieth that of man genes), as well as for the much higher fraction of coding Dna in their chromosomes. In addition to introns and exons, each gene is associated with regulatory Deoxyribonucleic acid sequences, which are responsible for ensuring that the factor is expressed at the proper level and time, and the proper type of cell. In humans, the regulatory sequences for a typical cistron are spread out over tens of thousands of nucleotide pairs. Every bit would be expected, these regulatory sequences are more compressed in organisms with compact genomes. Nosotros discuss in Chapter 7 how regulatory Deoxyribonucleic acid sequences piece of work.
Finally, the nucleotide sequence of the human genome has revealed that the critical information seems to be in an alarming state of disarray. Equally one commentator described our genome, "In some means it may resemble your garage/bedroom/fridge/life: highly individualistic, merely unkempt; trivial evidence of arrangement; much accumulated clutter (referred to by the uninitiated every bit 'junk'); virtually zippo always discarded; and the few patently valuable items indiscriminately, plainly carelessly, scattered throughout."
Comparisons Between the DNAs of Related Organisms Distinguish Conserved and Nonconserved Regions of Dna Sequence
A major obstacle in interpreting the nucleotide sequences of homo chromosomes is the fact that much of the sequence is probably unimportant. Moreover, the coding regions of the genome (the exons) are typically establish in short segments (average size about 145 nucleotide pairs) floating in a body of water of DNA whose verbal nucleotide sequence is of little consequence. This organisation makes it very difficult to identify all the exons in a stretch of DNA sequence; even harder is the conclusion of where a factor begins and ends and how many exons it spans. Accurate gene identification requires approaches that extract information from the inherently low signal-to-racket ratio of the human being genome, and we describe some of them in Chapter 8. Hither we hash out the most general approach, one that has the potential to identify not only coding sequences only also additional DNA sequences that are of import. It is based on the observation that sequences that have a function are conserved during development, whereas those without a function are free to mutate randomly. The strategy is therefore to compare the man sequence with that of the corresponding regions of a related genome, such every bit that of the mouse. Humans and mice are thought to have diverged from a common mammalian ancestor nigh 100 × ten6 years ago, which is long plenty for the majority of nucleotides in their genomes to have been changed past random mutational events. Consequently, the simply regions that will have remained closely similar in the ii genomes are those in which mutations would have impaired function and put the animals conveying them at a disadvantage, resulting in their emptying from the population past natural pick. Such closely similar regions are known as conserved regions. In general, conserved regions represent functionally important exons and regulatory sequences. In contrast, nonconserved regions represent DNA whose sequence is by and large not critical for part. By revealing in this manner the results of a very long natural "experiment," comparative Deoxyribonucleic acid sequencing studies highlight the most interesting regions in genomes.
Comparative studies of this kind take revealed non only that mice and humans share nearly of the same genes, but also that big blocks of the mouse and human genomes contain these genes in the same gild, a characteristic called conserved synteny (Figure 4-18). Conserved synteny can also exist revealed by chromosome painting, and this technique has been used to reconstruct the evolutionary history of our own chromosomes past comparing them with those from other mammals (Figure 4-xix).
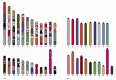
Figure four-18
Conserved synteny between the man and mouse genomes. Regions from different mouse chromosomes (indicated by the colors of each mouse in B) show conserved synteny (gene social club) with the indicated regions of the man genome (A). For example the genes (more...)

Figure 4-19
A proposed evolutionary history of human being chromosome 3 and its relatives in other mammals. (A) At the lower left is the social club of chromosome 3 segments hypothesized to exist present on a chromosome of a mammalian ancestor. Along the superlative are the patterns of (more...)
Chromosomes Exist in Different States Throughout the Life of a Cell
Nosotros accept seen how genes are arranged in chromosomes, merely to form a functional chromosome, a DNA molecule must exist able to do more than simply carry genes: it must be able to replicate, and the replicated copies must be separated and reliably partitioned into girl cells at each cell division. This process occurs through an ordered series of stages, collectively known as the cell cycle . The cell cycle is briefly summarized in Figure 4-xx, and discussed in detail in Chapter 17. Only ii of the stages of the wheel concern us in this chapter. During interphase chromosomes are replicated, and during mitosis they become highly condensed and so are separated and distributed to the two daughter nuclei. The highly condensed chromosomes in a dividing cell are known as mitotic chromosomes. This is the form in which chromosomes are most hands visualized; in fact, all the images of chromosomes shown so far in the affiliate are of chromosomes in mitosis. This condensed land is important in allowing the duplicated chromosomes to be separated past the mitotic spindle during prison cell division, as discussed in Affiliate 18.

Figure four-20
A simplified view of the eucaryotic cell cycle. During interphase, the prison cell is actively expressing its genes and is therefore synthesizing proteins. Also, during interphase and before jail cell division, the Dna is replicated and the chromosomes are duplicated. (more...)
During the portions of the cell cycle when the cell is not dividing, the chromosomes are extended and much of their chromatin exists as long, thin tangled threads in the nucleus so that individual chromosomes cannot be easily distinguished (Effigy 4-21). We refer to chromosomes in this extended state as interphase chromosomes.

Figure four-21
A comparison of extended interphase chromatin with the chromatin in a mitotic chromosome. (A) An electron micrograph showing an enormous tangle of chromatin spilling out of a lysed interphase nucleus. (B) A scanning electron micrograph of a mitotic chromosome: (more...)
Each Deoxyribonucleic acid Molecule That Forms a Linear Chromosome Must Contain a Centromere, Two Telomeres, and Replication Origins
A chromosome operates equally a distinct structural unit: for a copy to be passed on to each daughter prison cell at partitioning, each chromosome must be able to replicate, and the newly replicated copies must afterwards exist separated and partitioned correctly into the two daughter cells. These basic functions are controlled by three types of specialized nucleotide sequence in the Deoxyribonucleic acid, each of which binds specific proteins that guide the machinery that replicates and segregates chromosomes (Figure four-22).
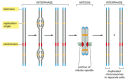
Figure 4-22
The 3 DNA sequences required to produce a eucaryotic chromosome that can be replicated and then segregated at mitosis. Each chromosome has multiple origins of replication, one centromere, and two telomeres. Shown here is the sequence of events a typical (more than...)
Experiments in yeasts, whose chromosomes are relatively small and easy to manipulate, take identified the minimal DNA sequence elements responsible for each of these functions. One blazon of nucleotide sequence acts equally a Dna replication origin, the location at which duplication of the Dna begins. Eucaryotic chromosomes comprise many origins of replication to ensure that the entire chromosome tin be replicated rapidly, equally discussed in detail in Chapter 5.
Later replication, the 2 daughter chromosomes remain fastened to ane another and, as the jail cell bike proceeds, are condensed further to produce mitotic chromosomes. The presence of a 2d specialized Dna sequence, called a centromere, allows i copy of each duplicated and condensed chromosome to be pulled into each daughter cell when a cell divides. A protein circuitous chosen a kinetochore forms at the centromere and attaches the duplicated chromosomes to the mitotic spindle, assuasive them to be pulled apart (discussed in Chapter 18).
The third specialized Deoxyribonucleic acid sequence forms telomeres, the ends of a chromosome. Telomeres contain repeated nucleotide sequences that enable the ends of chromosomes to be efficiently replicated. Telomeres also perform another part: the repeated telomere DNA sequences, together with the regions bordering them, form structures that protect the cease of the chromosome from being recognized by the cell as a broken DNA molecule in demand of repair. Nosotros discuss this type of repair and the other features of telomeres in Chapter five.
In yeast cells, the 3 types of sequences required to propagate a chromosome are relatively short (typically less than thousand base pairs each) and therefore use merely a tiny fraction of the information-carrying capacity of a chromosome. Although telomere sequences are fairly uncomplicated and short in all eucaryotes, the DNA sequences that specify centromeres and replication origins in more complex organisms are much longer than their yeast counterparts. For case, experiments suggest that human centromeres may contain upwards to 100,000 nucleotide pairs. It has been proposed that human centromeres may not fifty-fifty require a stretch of DNA with a defined nucleotide sequence; instead, they may simply create a large, regularly repeating protein-nucleic acid construction. We return to this event at the end of the affiliate when we discuss in more full general terms the proteins that, forth with Deoxyribonucleic acid, make up chromosomes.
Dna Molecules Are Highly Condensed in Chromosomes
All eucaryotic organisms accept elaborate ways of packaging DNA into chromosomes. Call up from earlier in this affiliate that human being chromosome 22 contains about 48 million nucleotide pairs. Stretched out end to end, its DNA would extend almost ane.5 cm. Yet, when it exists equally a mitotic chromosome, chromosome 22 measures only about 2 μm in length (see Figures iv-10 and 4-11), giving an end-to-finish compaction ratio of almost x,000-fold. This remarkable feat of compression is performed past proteins that successively coil and fold the Deoxyribonucleic acid into higher and higher levels of organization. Although less condensed than mitotic chromosomes, the Dna of interphase chromosomes is nevertheless tightly packed, with an overall compaction ratio of approximately 1000-fold. In the next sections we discuss the specialized proteins that make the pinch possible.
In reading these sections it is important to proceed in mind that chromosome construction is dynamic. Not merely do chromosomes globally condense in accord with the cell bike, but different regions of the interphase chromosomes condense and decondense every bit the cells gain access to specific DNA sequences for gene expression, DNA repair, and replication. The packaging of chromosomes must therefore be accomplished in a fashion that allows rapid localized, on-need access to the DNA.
Nucleosomes Are the Basic Unit of Eucaryotic Chromosome Structure
The proteins that demark to the Dna to class eucaryotic chromosomes are traditionally divided into two full general classes: the histones and the nonhistone chromosomal proteins. The complex of both classes of protein with the nuclear DNA of eucaryotic cells is known as chromatin. Histones are present in such enormous quantities in the jail cell (about 60 million molecules of each type per human cell) that their total mass in chromatin is nigh equal to that of the DNA.
Histones are responsible for the first and most basic level of chromosome organisation, the nucleosome, which was discovered in 1974. When interphase nuclei are broken open up very gently and their contents examined under the electron microscope, almost of the chromatin is in the form of a cobweb with a bore of about 30 nm (Figure 4-23A). If this chromatin is subjected to treatments that cause it to unfold partially, it can exist seen nether the electron microscope every bit a series of "chaplet on a cord" (Figure 4-23B). The string is DNA, and each dewdrop is a "nucleosome cadre particle" that consists of DNA wound effectually a protein core formed from histones. The beads on a cord represent the first level of chromosomal DNA packing.

Figure iv-23
Nucleosomes as seen in the electron microscope. (A) Chromatin isolated direct from an interphase nucleus appears in the electron microscope as a thread 30 nm thick. (B) This electron micrograph shows a length of chromatin that has been experimentally (more than...)
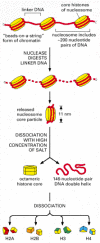
The structural arrangement of nucleosomes was adamant after first isolating them from unfolded chromatin by digestion with item enzymes (called nucleases) that intermission downwardly DNA past cut between the nucleosomes. After digestion for a brusk menstruation, the exposed DNA between the nucleosome core particles, the linker Deoxyribonucleic acid, is degraded. Each private nucleosome core particle consists of a circuitous of viii histone proteins—two molecules each of histones H2A, H2B, H3, and H4—and double-stranded Deoxyribonucleic acid that is 146 nucleotide pairs long. The histone octamer forms a protein core around which the double-stranded Deoxyribonucleic acid is wound (Figure 4-24).
Effigy 4-24
Structural organization of the nucleosome. A nucleosome contains a protein core made of eight histone molecules. Equally indicated, the nucleosome cadre particle is released from chromatin past digestion of the linker Deoxyribonucleic acid with a nuclease, an enzyme that breaks (more than...)
Each nucleosome core particle is separated from the next past a region of linker DNA, which can vary in length from a few nucleotide pairs up to about 80. (The term nucleosome technically refers to a nucleosome core particle plus 1 of its adjacent DNA linkers, but information technology is frequently used synonymously with nucleosome core particle.) On boilerplate, therefore, nucleosomes echo at intervals of about 200 nucleotide pairs. For case, a diploid homo prison cell with 6.four × ten9 nucleotide pairs contains approximately 30 million nucleosomes. The formation of nucleosomes converts a Deoxyribonucleic acid molecule into a chromatin thread nigh ane-tertiary of its initial length, and this provides the beginning level of Deoxyribonucleic acid packing.
The Structure of the Nucleosome Core Particle Reveals How Dna Is Packaged
The loftier-resolution structure of a nucleosome core particle, solved in 1997, revealed a disc-shaped histone core around which the Dna was tightly wrapped 1.65 turns in a left-handed coil (Effigy 4-25). All four of the histones that brand up the core of the nucleosome are relatively small proteins (102–135 amino acids), and they share a structural motif, known equally the histone fold, formed from three α helices connected past two loops (Effigy four-26). In assembling a nucleosome, the histone folds first bind to each other to form H3–H4 and H2A-H2B dimers, and the H3–H4 dimers combine to course tetramers. An H3–H4 tetramer so farther combines with two H2A-H2B dimers to class the compact octamer core, effectually which the Dna is wound (Figure 4-27).

Figure 4-25
The structure of a nucleosome core particle, every bit determined by x-ray diffraction analyses of crystals. Each histone is colored according to the scheme of Effigy iv-24, with the Dna double helix in light greyness. (Reprinted by permission from Yard. Luger et al., (more than...)

Figure 4-26
The overall structural organization of the core histones. (A) Each of the core histones contains an N-terminal tail, which is subject area to several forms of covalent modification, and a histone fold region, as indicated. (B) The structure of the histone (more...)
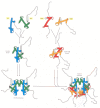
Figure 4-27
The assembly of a histone octamer. The histone H3–H4 dimer and the H2A-H2B dimer are formed from the handshake interaction. An H3-H4 tetramer forms the scaffold of the octamer onto which 2 H2A-H2B dimers are added, to complete the assembly. (more...)
The interface between DNA and histone is extensive: 142 hydrogen bonds are formed betwixt Deoxyribonucleic acid and the histone core in each nucleosome. Nearly one-half of these bonds course between the amino acid courage of the histones and the phosphodiester backbone of the DNA. Numerous hydrophobic interactions and salt linkages also concord Deoxyribonucleic acid and poly peptide together in the nucleosome. For instance, all the core histones are rich in lysine and arginine (two amino acids with basic side chains), and their positive charges can finer neutralize the negatively charged DNA backbone. These numerous interactions explain in part why DNA of virtually any sequence can be bound on a histone octamer cadre. The path of the Deoxyribonucleic acid around the histone core is not smooth; rather, several kinks are seen in the DNA, as expected from the nonuniform surface of the core.
In addition to its histone fold, each of the cadre histones has a long N-terminal amino acid "tail", which extends out from the Deoxyribonucleic acid-histone core (see Figure 4-27). These histone tails are subject to several different types of covalent modifications, which control many aspects of chromatin structure. We discuss these issues later in the chapter.
Equally might be expected from their fundamental role in Deoxyribonucleic acid packaging, the histones are among the almost highly conserved eucaryotic proteins. For case, the amino acid sequence of histone H4 from a pea and a cow differ at only at 2 of the 102 positions. This strong evolutionary conservation suggests that the functions of histones involve virtually all of their amino acids, so that a alter in any position is deleterious to the jail cell. This suggestion has been tested directly in yeast cells, in which information technology is possible to mutate a given histone factor in vitro and innovate it into the yeast genome in place of the normal cistron. As might be expected, virtually changes in histone sequences are lethal; the few that are not lethal cause changes in the normal pattern of gene expression, as well equally other abnormalities.
Despite the loftier conservation of the cadre histones, many eucaryotic organisms too produce specialized variant cadre histones that differ in amino acrid sequence from the main ones. For example, the sea urchin has five histone H2A variants, each of which is expressed at a different time during development. Information technology is thought that nucleosomes that accept incorporated these variant histones differ in stability from regular nucleosomes, and they may be peculiarly well suited for the high rates of DNA transcription and DNA replication that occur during these early stages of development.
The Positioning of Nucleosomes on DNA Is Determined by Both Dna Flexibility and Other Dna-bound Proteins
Although nearly every DNA sequence can, in principle, exist folded into a nucleosome, the spacing of nucleosomes in the cell tin can be irregular. Two primary influences determine where nucleosomes course in the DNA. Ane is the difficulty of bending the DNA double helix into two tight turns around the outside of the histone octamer, a process that requires substantial compression of the pocket-size groove of the DNA helix. Because A-T-rich sequences in the small-scale groove are easier to compress than G-C-rich sequences, each histone octamer tends to position itself on the Deoxyribonucleic acid so as to maximize A-T-rich minor grooves on the inside of the Deoxyribonucleic acid coil (Figure 4-28). Thus, a segment of Deoxyribonucleic acid that contains short A-T-rich sequences spaced by an integral number of Dna turns is easier to bend around the nucleosome than a segment of Deoxyribonucleic acid defective this feature. In improver, because the Deoxyribonucleic acid in a nucleosome is kinked in several places, the ability of a given nucleotide sequence to accommodate this deformation can also influence the position of DNA on the nucleosome.
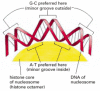
Figure 4-28
The bending of Deoxyribonucleic acid in a nucleosome. The DNA helix makes 1.65 tight turns around the histone octamer. This diagram is drawn approximately to scale, illustrating how the small-scale groove is compressed on the inside of the turn. Attributable to sure structural features (more...)
These features of DNA probably explain some hitting, but unusual, cases of very precise positioning of nucleosomes along a stretch of DNA. For most of the DNA sequences found in chromosomes, however, there is no strongly preferred nucleosome-binding site; a nucleosome can occupy any one of a number of positions relative to the DNA sequence.
The second, and probably about important, influence on nucleosome positioning is the presence of other tightly bound proteins on the Deoxyribonucleic acid. Some jump proteins favor the formation of a nucleosome adjacent to them. Others create obstacles that force the nucleosomes to get together at positions betwixt them. Finally, some proteins can bind tightly to DNA fifty-fifty when their DNA-binding site is part of a nucleosome. The verbal positions of nucleosomes along a stretch of Deoxyribonucleic acid therefore depend on factors that include the Deoxyribonucleic acid sequence and the presence and nature of other proteins bound to the Deoxyribonucleic acid. Moreover, as we see below, the arrangement of nucleosomes on Deoxyribonucleic acid is highly dynamic, changing rapidly according to the needs of the cell.
Nucleosomes Are Usually Packed Together into a Compact Chromatin Fiber
Although long strings of nucleosomes grade on most chromosomal Dna, chromatin in a living prison cell probably rarely adopts the extended "chaplet on a cord" grade. Instead, the nucleosomes are packed on top of one another, generating regular arrays in which the Deoxyribonucleic acid is even more highly condensed. Thus, when nuclei are very gently lysed onto an electron microscope filigree, most of the chromatin is seen to be in the form of a fiber with a bore of about thirty nm, which is considerably wider than chromatin in the "beads on a cord" course (see Figure four-23).
Several models have been proposed to explain how nucleosomes are packed in the 30-nm chromatin fiber; the one near consequent with the available information is a serial of structural variations known collectively every bit the Zigzag model (Figure 4-29). In reality, the xxx-nm structure constitute in chromosomes is probably a fluid mosaic of the different zigzag variations. Nosotros saw earlier that the linker DNA that connects next nucleosomes tin can vary in length; these differences in linker length probably innovate further local perturbations into the zigzag structure. Finally, the presence of other DNA-binding proteins and Dna sequence that are difficult to fold into nucleosomes punctuate the 30-nm fiber with irregular features (Figure 4-30).

Figure 4-29
Variations on the Zigzag model for the xxx-nm chromatin fiber. (A and B) Electron microscopic bear witness for the top and lesser-left model structures depicted in (C). (C) Zigzag variations. An interconversion between these three variations is proposed to (more than...)
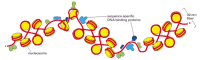
Figure 4-30
Irregularities in the 30-nm fiber. This schematic view of the xxx-nm cobweb illustrates its suspension past sequence-specific DNA-binding proteins. How these proteins bind tightly to DNA is explained in Chapter 7. The interruptions in the 30-nm fiber may (more...)
Several mechanisms probably act together to grade the xxx-nm cobweb from a linear string of nucleosomes. First, an additional histone, chosen histone H1, is involved in this process. H1 is larger than the core histones and is considerably less well conserved. In fact, the cells of most eucaryotic organisms make several histone H1 proteins of related just quite distinct amino acid sequences. A single histone H1 molecule binds to each nucleosome, contacting both Deoxyribonucleic acid and protein, and changing the path of the Dna equally information technology exits from the nucleosome. Although it is not understood in detail how H1 pulls nucleosomes together into the 30-nm fiber, a alter in the exit path in DNA seems crucial for compacting nucleosomal Deoxyribonucleic acid so that it interlocks to class the 30-nm fiber (Figure 4-31).
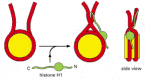
Figure four-31
A speculative model for how histone H1 could change the path of Dna as it exits from the nucleosome. Histone H1 (green) consists of a globular cadre and ii extended tails. Part of the upshot of H1 on the compaction of nucleosome arrangement may outcome (more...)
A second machinery for forming the xxx-nm fiber probably involves the tails of the cadre histones, which, as we saw higher up, extend from the nucleosome. It is idea that these tails may help attach 1 nucleosome to another—thereby assuasive a cord of them, with the help of histone H1, to condense into the 30-nm fiber (Figure four-32).

Figure 4-32
A speculative model for histone tails in the formation of the 30-nm cobweb. (A) The guess exit points of the 8 histone tails, 4 from each histone subunit, that extend from each nucleosome. In the high-resolution structure of the nucleosome (more...)
ATP-driven Chromatin Remodeling Machines Change Nucleosome Structure
For many years biologists thought that, one time formed in a particular position on DNA, a nucleosome remained stock-still in place because of the tight association between the core histones and DNA. But it has recently been discovered that eucaryotic cells incorporate chromatin remodeling complexes, protein machines that use the free energy of ATP hydrolysis to change the structure of nucleosomes temporarily and so that DNA becomes less tightly spring to the histone core. The remodeled country may result from motion of the H2A-H2B dimers in the nucleosome core; the H3–H4 tetramer is specially stable and would exist hard to rearrange (run into Effigy four-27).
The remodeling of nucleosome structure has 2 of import consequences. First, it permits ready access to nucleosomal Deoxyribonucleic acid by other proteins in the cell, particularly those involved in gene expression, DNA replication, and repair. Even afterwards the remodeling circuitous has dissociated, the nucleosome can remain in a "remodeled state" that contains Deoxyribonucleic acid and the total complement of histones—but ane in which the DNA-histone contacts have been loosened; merely gradually does this remodeled country revert to that of a standard nucleosome. Second, remodeling complexes can catalyze changes in the positions of nucleosomes along Deoxyribonucleic acid (Figure 4-33); some tin can fifty-fifty transfer a histone core from one DNA molecule to some other.

Figure 4-33
Model for the mechanism of some chromatin remodeling complexes. In the absence of remodeling complexes, the interconversion betwixt the three nucleosomal states shown is very slow considering of a high activation free energy barrier. Using ATP hydrolysis, chromatin-remodeling (more...)
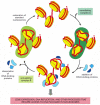
Cells have several different chromatin remodeling complexes that differ subtly in their properties. Most are large protein complexes that can contain more than ten subunits. It is likely that they are used whenever a eucaryotic jail cell needs direct access to nucleosome DNA for factor expression, Dna replication, or Dna repair. Unlike remodeling complexes may take features specialized for each of these roles. It is thought that the primary role of some remodeling complexes is to allow admission to nucleosomal Dna, whereas that of others is to re-grade nucleosomes when admission to Dna is no longer required (Effigy 4-34).
Effigy 4-34
A cyclic mechanism for nucleosome disruption and re-formation. According to this model, different chromatin remodeling complexes disrupt and re-grade nucleosomes, although, in principle, the aforementioned complex might catalyze both reactions. The DNA-bounden proteins (more...)
Chromatin remodeling complexes are carefully controlled by the cell. We shall see in Affiliate vii that, when genes are turned on and off, these complexes can be brought to specific regions of DNA where they act locally to influence chromatin structure. During mitosis, at least some of the chromatin-remodeling complexes are inactivated by phosphorylation. This may assistance the tightly packaged mitotic chromosomes maintain their construction.
Covalent Modification of the Histone Tails Can Profoundly Affect Chromatin
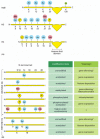
The Due north-terminal tails of each of the four core histones are highly conserved in their sequence, and perform crucial functions in regulating chromatin structure. Each tail is subject to several types of covalent modifications, including acetylation of lysines, methylation of lysines, and phosphorylation of serines (Figure 4-35A). Histones are synthesized in the cytosol and so assembled into nucleosomes. Some of the modifications of histone tails occur just after their synthesis, but before their assembly. The modifications that business united states of america, however, take place one time the nucleosome has been assembled. These nucleosome modifications are added and removed by enzymes that reside in the nucleus; for example, acetyl groups are added to the histone tails past histone acetyl transferases (HATs) and taken off by histone deacetylases (HDACs).
Effigy 4-35
Covalent modification of core histone tails. (A) Known modifications of the four histone cadre proteins are indicated: Me = methyl group, Ac = acetyl group, P = phosphate, u = ubiquitin. Note that some positions (e.m., lysine 9 of H3) can be modified in (more...)
The various modifications of the histone tails accept several important consequences. Although modifications of the tails take niggling directly upshot on the stability of an private nucleosome, they seem to bear upon the stability of the 30-nm chromatin fiber and of the higher-lodge structures discussed beneath. For example, histone acetylation tends to destabilize chromatin structure, perhaps in function because adding an acetyl group removes the positive accuse from the lysine, thereby making it more difficult for histones to neutralize the charges on DNA as chromatin is compacted. However, the most profound effect of modified histone tails is their ability to attract specific proteins to a stretch of chromatin that has been appropriately modified. Depending on the precise tail modifications, these boosted proteins tin either crusade further compaction of the chromatin or can facilitate access to the Deoxyribonucleic acid. If combinations of modifications are taken into account, the number of possible distinct markings for each histone tail is very large. Thus, it has been proposed that, through covalent modification of the histone tails, a given stretch of chromatin can convey a particular meaning to the cell (Figure iv-35B). For case, one type of marking could signal that the stretch of chromatin has been newly replicated, and another could indicate that gene expression should non take identify. According to this idea, each different mark would attract those proteins that would so execute the appropriate functions. Because the histone tails are extended, and are therefore probably attainable fifty-fifty when chromatin is condensed, they provide an especially apt format for such messages.
As with chromatin remodeling complexes, the enzymes that modify (and remove modifications from) histone tails are usually multisubunit proteins, and they are tightly regulated. They are brought to a particular region of chromatin by other cues, peculiarly past sequence-specific Deoxyribonucleic acid-bounden proteins. We can thus imagine how cycles of histone tail modification and demodification can allow chromatin structure to exist dynamic—locally compacting and decompacting information technology, and, in improver, attracting other proteins specific for each modification country. It is likely that histone-modifying enzymes and chromatin remodeling complexes work in concert to condense and recondense stretches of chromatin; for example, testify suggests that a item modification of the histone tail attracts a particular type of remodeling complex. Moreover, some chromatin remodeling complexes incorporate histone modification enzymes every bit subunits, directly connecting the two processes.
Summary
A gene is a nucleotide sequence in a DNA molecule that acts as a functional unit for the production of a protein, a structural RNA, or a catalytic RNA molecule. In eucaryotes, protein-coding genes are normally equanimous of a string of alternating introns and exons. A chromosome is formed from a single, enormously long DNA molecule that contains a linear assortment of many genes. The man genome contains 3.ii × 10 ix DNA nucleotide pairs, divided betwixt 22 different autosomes and 2 sex chromosomes. Only a small percentage of this DNA codes for proteins or structural and catalytic RNAs. A chromosomal Deoxyribonucleic acid molecule also contains three other types of functionally of import nucleotide sequences: replication origins and telomeres allow the DNA molecule to be completely replicated, while a centromere attaches the daughter DNA molecules to the mitotic spindle, ensuring their authentic segregation to daughter cells during the M phase of the cell cycle.
The DNA in eucaryotes is tightly jump to an equal mass of histones, which form a repeating array of Deoxyribonucleic acid-protein particles called nucleosomes. The nucleosome is composed of an octameric core of histone proteins around which the DNA double helix is wrapped. Despite irregularities in the positioning of nucleosomes along DNA, nucleosomes are usually packed together (with the aid of histone H1 molecules) into quasi-regular arrays to class a thirty-nm fiber. Despite the high degree of compaction in chromatin, its structure must be highly dynamic to let the prison cell access to the Dna. 2 general strategies for reversibly changing local chromatin structures are important for this purpose: ATP-driven chromatin remodeling complexes, and an enzymatically catalyzed covalent modification of the N-concluding tails of the four cadre histones.
Source: https://www.ncbi.nlm.nih.gov/books/NBK26834/
Posted by: madsonvalln1946.blogspot.com

0 Response to "What Does Condensed Dna Makeup A Chromosome"
Post a Comment